TAMBIS
[ Home ] [ Project Details
] [ Personnel ] [ Publications
]
Aims
TAMBIS aims to provide transparent information retrieval and filtering
from biological information services by building a homogenising layer on
top of the different sources. This layer uses a mediator and many
source wrappers to create the illusion of one all encompassing data
source.
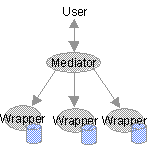 The mediator is a kind of information broker. It uses a conceptual knowledge
base of molecular biology, expressed in the Description Logic GRAIL, to
The mediator is a kind of information broker. It uses a conceptual knowledge
base of molecular biology, expressed in the Description Logic GRAIL, to
-
describe the universal model
-
help users form queries against this universal model expressed in the modelling
language
-
mediate between the various sources to translate the mediators model to
the sources models
The wrappers create the illusion of a common query language for each information
resource. The current wrapper language is CPL.
The mediator takes conceptual and source-independent GRAIL queries and
rewrites them to concrete and source-dependent CPL queries, which are despatched
to the sources.
On the way to building TAMBIS we have developed a number of other generally
useful things including:
-
a biological ontology
-
a questionnaire on bioinformaticians requirements
-
a benchmark query set
Biological Terminology Concept Server
The BioCon Knowledge Base is an ontology of biological concepts.
The Terminology Server provides a number of services for terminological
or concept models such as what can I say about concept X? or is
concept Y coherent? or what are the immediate parents and/or children
of concept Z?
Together they form a Biological Terminology Concept Server used
by TAMBIS.
The BioCon Knowledge Base is used to:
-
describe the metadata of the underlying data sources, representing an over-arching
universal schema;
-
form queries expressed in the modelling language;
-
drive a GUI user interface for query formulation as users are not prepared
to handle complex query languages;
-
mediate between the various sources to translate the mediators model to
the sources models. Mediation exploits the BioCon lattice of concepts
to assist in the identification and resolution of equivalences or near
equivalences in information sources.
System architecture
The architecture of the TAMBIS system is shown in the figure below.
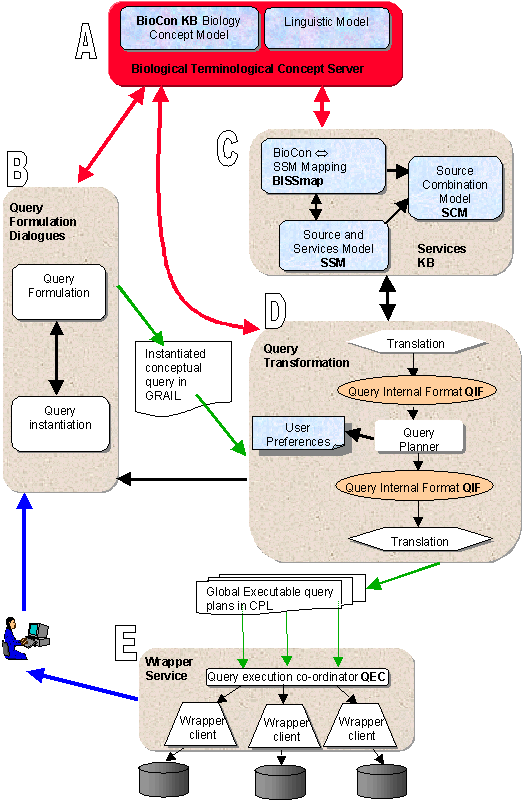
The main components are:
A. An ontology of biological terms in GRAIL
B. A knowledge-driven query formulation interface
C. A services model linking the biological ontology with the
source schemas
D. A query transformation rewriting process
E. A wrapper service dealing with external sources
The User Interface
Queries are constructed using a graphical query interface which allows
users to piece together queries and insulates them from the syntax of the
underlying description logic model.
Two applets demonstrating the TAMBIS system are accessible.
The two applets has the same interface but with different
capability.
There are three videos demonstrating how queries can be built and
processed using the applets.
[ Home ] [ Project
Details ] [ Personnel ] [ Publications
]


